|
Make sure to use a decimal in your number or your command will not work properly (ie: use "wireframe 1.0", not "wireframe 1"). You can also use a number to size your wireframe lines to an exact diameter (in angstroms). But you can scale the lines up by including a percent to the end of your wireframe command. 
To apply Wireframe Format, enter the command.īy default, wireframe lines are very thin. By default, all structures opened in the JUDE environment will initially be displayed in Wireframe Format. Wireframe Format displays all of the covalent bonds in a protein structure as thin lines, and is a great way to see how atoms are connected. There are three main Display Formats that can be applied to a protein structure using the Jmol commands entered into the console. The Jmol commands being demonstrated will always appear as Orange Text throughout this training guide. Note: A Jmol Console has also been added to the bottom right corner of this web page so that you can practice any Jmol Commands being discussed without leaving the page. Once a Jmol command has been typed into the console, you can either click the "Execute" button or press the "Enter" key on your keyboard to submit the command. In the bottom right corner of the Jmol User Design Environment is the Jmol Console, which will be used to input text commands that can alter your Jmol display. 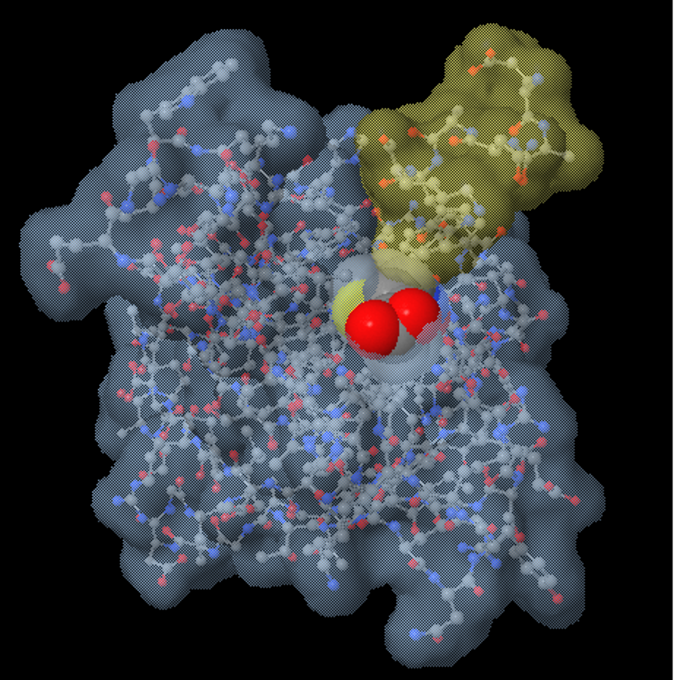
This structure file includes one protein strand and two DNA strands. You can load this structure in to the JUDE webpage using the PDB ID "1ZAA". Note: For the remainder of this Jmol Training Guide, we will be using the Zinc Finger protein as our example protein structure (based on 1ZAA.pdb). In this section of the Jmol Training Guide, you will learn how to first Explore a Protein Structure using the Display Format and Color Format commands, and then how to Focus in on the Backbone, as shown in the image above and to the right. It is hard to see the path of the amino acid strand(s) or locate individual amino acid sidechains that may be key to the Protein Story you want your model design to tell. While visually impressive, this default display can also be overwhelming. When a Protein Structure (.pdb) File is first opened in the Jmol User Design Environment (JUDE), it will display every atom colored by element (carbon=gray, oxygen=red, nitrogen=blue) with thin lines representing covalent bonds, as shown in the image below and to the left. Proteins are long strands of amino acids that fold into complex 3-dimensional shapes following basic principles of chemistry. 
Part 2 - Simplifying Your Protein The CBM Jmol Training Guide Introduction
0 Comments
Leave a Reply. |
 RSS Feed
RSS Feed
